A Pan-Tumor Assay
for Solid Tumors
Pan-Tumor Assay for Solid Tumors
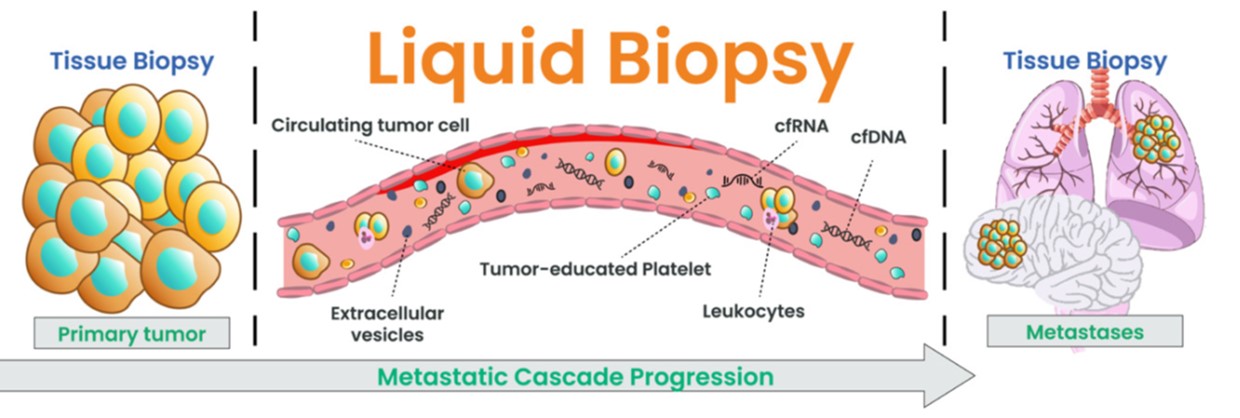
GTC’s Liquid Trace Solid Tumor is a highly sensitive pan-cancer test that evaluates cell-free RNA and DNA (cfRNA and cfDNA), providing highly informative data that can be used for diagnoses, evaluating the host immune response, and identifying biomarkers for predicting responses to various therapies.
Furthermore, Liquid Trace Solid Tumor may provide additional information on changes not detected with tissue biopsies, including germline mutations and mutations in the subclones not present in the tissue sample (heterogeneity).
Examples of types of solid tumors Liquid Trace can detect:
- Lung
- Brain
- Breast
- Thyroid
- Colon
- Oropharyngeal tumors
- Pancreatic
- Ovarian
- Prostate
Many conventional liquid biopsy tests are dependent solely on cfDNA analysis, which presents multiple challenges. These include variations in DNA shedding between tumors as well as low sensitivity (especially in early-stage cancer), difficulty in detecting fusion genes (i.e., chromosomal translocations leading to the expression of chimeric mRNA from two genes), and inability to detect the numerous biological processes that modify RNA expression levels, such as alternative splicing, stability, and allele-specific methylation. The latter limitation is critically important as recent studies have shown that RNA testing provides another level of biological information regarding the tumor and its microenvironment.
-TMB evaluation is now part of Liquid Trace Solid Tumor
The Benefits of cfRNA
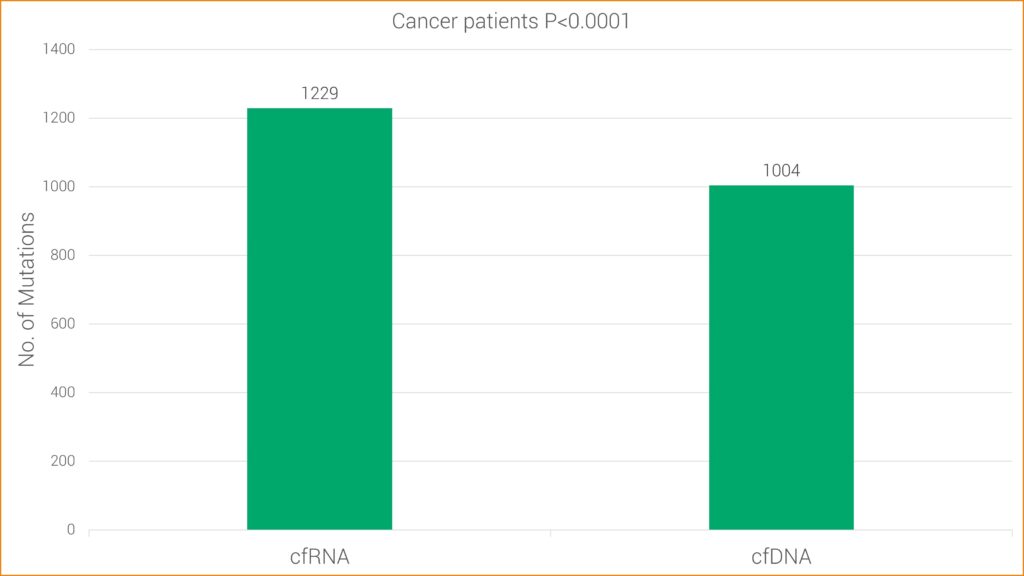
Studies have found RNA sequencing to be more sensitive for some types of mutations, likely because cancer cells typically contain one copy of mutated DNA but numerous copies of RNA. This research is consistent with GTC’s findings that cfRNA has increased sensitivity over cfDNA alone. More specifically, cfRNA allows GTC’s Liquid Trace to detect mutations and fusions in hematologic and solid tumor samples that may be undetected with conventional cfDNA testing.
T-cell and B-cell Clonality Detection: The detection of T- and B-cell clonality is important because it can help diagnose and monitor certain types of malignancies. When malignant transformation occurs in a T- or B-cell, the cells can undergo uncontrolled clonal expansion, resulting in the accumulation of many cells with the identical T- or B-cell receptor.
HLA Class I Genotyping: Useful when identifying patients who may be candidates for immunotherapy.
Epstein-Barr Virus (EBV): Important for diagnosis and classification of lymphoid neoplasms and some solid tumors.
Human Papillomavirus (HPV): Chronic infection with high-risk HPV subtypes is associated with increased risk of anogenital and oropharyngeal cancer. Detection of HPV mRNA suggests active (productive) infection.
Torque Teno Virus (TTV): This virus was first discovered in a patient with non-A-E hepatitis and is now regarded as a part of the human virome. In general, TTV does not cause pathology in immunocompetent individuals. This virus is considered as a marker of the degree of immune competence in patients with immunological impairment and inflammatory disorders. High TTV load is associated with increased risk of infection. In patients with organ transplant, low TTV load is associated with an increased risk of rejection.
Sensitivity is 0.1 to 0.01 for non-hot spot, 0.01 to 0.001 for hotspot and <0.001 for tumor informed or prior Hx.
For DNA, QNS is rare (<0.1%), but it is higher for RNA (Good DNA results but poor RNA results. Of course, if we receive 3 ml of plasma (6 ml blood), the sample is QNS for performing RNA testing.
VAF (Variant Allele Frequency) value: This value is used to monitor the disease in liquid bx. The high the VAF means higher tumor load. Patients showing reduction in VAF after treatment means they are doing better.
GTC uses AI in every step of our analysis and it makes a difference in helping make a new discovery daily that improves patient care.
Once the data is offloaded from the sequencer, our AI:
- Assists with mutation analysis, identifying non-mutations and artifacts
- Compares various data sets to explore disease biology
- Provides support for clinical decision-making and classification of the disease
- It helps with matching patients to therapeutics and presents clinical trial options
- Aggregates data for report generation and simplifies the results so they are easily understood
Case Study: Prostate Cancer
Background
Prostate cancer is one of the most common solid tumors among men. Multiple therapies have been introduced to improve survival and symptom control. Analysis of tumor cfDNA and cfRNA in the blood, using liquid biopsies, has become an important tool in the management of prostate cancer. In localized disease, it can distinguish between low-and high-grade cancers and can guide the decision to proceed with or defer tissue biopsy. In advanced tumor states, liquid biopsy has a prognostic value and has been used in clinical trials to assess response.
Clinical History
- 75-year-old male with a history of prostate cancer presenting for monitoring
Molecular Profiling Findings
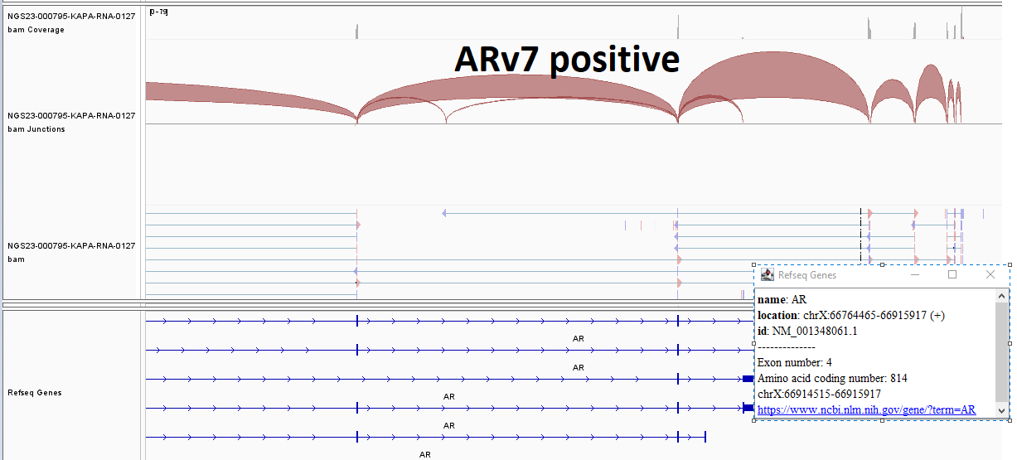
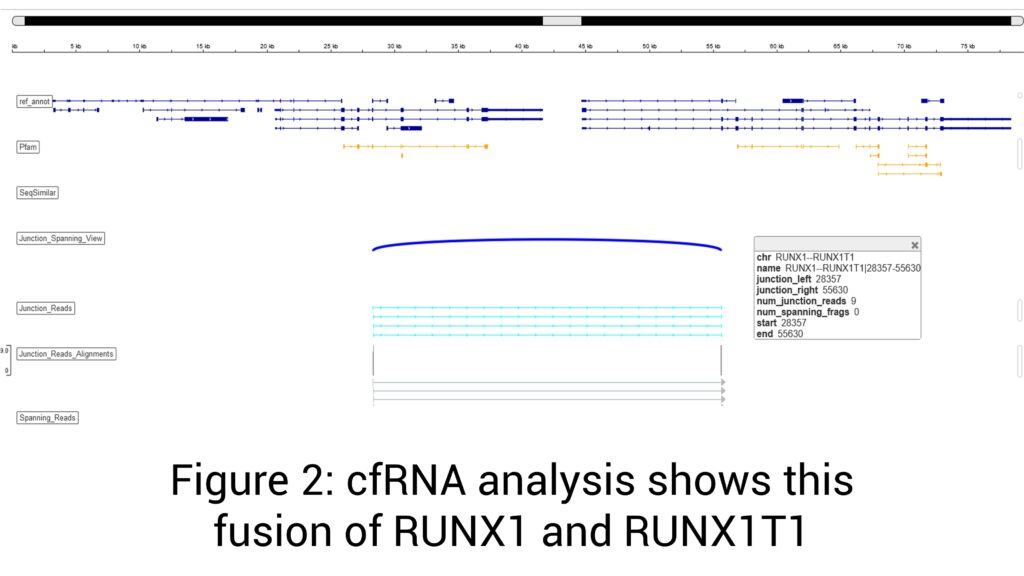
Androgen receptor splice variant 7 (AR-V7) detected (Figure 1)
t(21;21)(q22;q22) ERG-TMPRSS2 mRNA fusion
Mutations in TP53, CDK12, and GATA2 genes
Chromosomal structural analysis shows +7, +8, -11, +12, -13, -14, and others
Increased prostate-specific antigen (PSA) mRNA
Increased keratin 19 mRNA
No evidence of germline BRCA1/2, BARD1, BRIP1, CDK12, CHEK1/2, FANCL, PALB2, or RAD mutation
Discussion
Although serum PSA is used to monitor prostate cancer, PSA levels have often failed to precisely reflect disease burden and extent. Multiple therapies impact patient survival and symptoms without corresponding changes in serum PSA levels. As such, comprehensive analysis of cfDNA and cfRNA using liquid biopsies provides another level of biological information regarding the tumor and its microenvironment. This technique is simple, safe, and easily repeatable throughout the disease course and can serve as a prognostic and predictive biomarker as well as a ready tissue source for molecular profiling. In this specific case we were able to evaluate the patient’s mutation status for AR-V7 (at the RNA level) and homologous recombination repair genes. Men with AR-V7 expression have a shorter progression-free and overall survival when treated with enzalutamide or abiraterone, suggesting a possible means of predicting response to these therapies through cfDNA and cfRNA profiling.
Download Case Study Brochure (PDF)
References
- Siegel, R. L., Miller, K. D., & Jemal, A. (2019). authors. Cancer Statistics, 2019. CA Cancer J Clin, 69, 7-34
- Albitar, M., Zhang, H., Charifa, A., Ip, A., De Dios, I., Ma, W., ... & Goy, A. (2022). Cell-free RNA in liquid biopsy and biomarkers profiling of hematologic and solid tumors. Clin Chem Lab Med, 60, 1855-1867.
- Albitar, M., Zhang, H., Charifa, A., Ip, A., De Dios, I., Ma, W., ... & Goy, A. (2022). Combining cell-free RNA (cfRNA) with cell-free total nucleic acid (cfTNA) as a new paradigm for liquid biopsy. Abstract presented at: 2022 ASCO Annual Meeting; June 3-7, 2022; Chicago, IL.
Case Study: ESR1
Background
Circulating tumor DNA and RNA is circulating cell-free DNA/RNA (cfDNA/cfRNA) released by tumor cells in the blood. cfDNA and cfRNA can be detected in the plasma of patients with cancer, and their analysis represents a minimally invasive tool for detecting and monitoring key gene mutations and alterations. In breast cancer, detection of mutations in the estrogen receptor 1 (ESR1) and PIK3CA genes is very important, since specific targeted therapies have recently there have been developed for patients with mutations in these genes.
Clinical History
- 62-year-old female with relapsing (ER+/HER2 -) breast cancer
Molecular Profiling Findings
- Mutations in ESR1, PIK3CA, MAP3K1, PRKDC, KMT2C, MYC, ROS1, NOTCH2, and EP300 genes
Discussion
Endocrine therapy is the main treatment option for estrogen receptor-positive (ER+) breast cancer (BC). Compared with other clinical subtypes, ER+ BC patients usually have a more favorable prognosis. However, almost all ER+ BC patients eventually develop endocrine resistance and disease progression. Mutations in ESR1 play a key role in resistance to aromatase inhibitors, but tumors harboring these mutations may retain sensitivity to selective estrogen receptor degraders (SERD). Recently, elacestrant, an oral SERD, was approved for patients with ER+, HER2-negative, ESR1 mutant advanced or metastatic breast cancer. Detection of PIK3CA mutations is important as well, since alpelisib in combination with fulvestrant has been approved for patients with HR+, HER2-negative, PIK3CA mutant advanced or metastatic breast cancer whose disease has progressed after endocrine therapy. Detection of ESR1 and PIK3CA mutations could also have important clinical applications for the follow-up of these patients. In this case, the patient’s breast cancer tissue was sequenced initially, and treated was prescribed accordingly. ESR1 mutation was not detected in the diagnostic tissue sample. Later, the patient’s disease progressed and relapsed. A minimally invasive liquid biopsy revealed ESR1 mutation, consistent with endocrine resistance.
In summary, novel SERDS and PIK3CA inhibitors are emerging as new therapeutic options for metastatic breast cancer. The analysis of cfDNA and cfRNA allows for non-invasive monitoring of these mutations over time, providing clinicians with valuable information about treatment response, disease progression, and the emergence of resistance. This approach reduces the need for invasive tissue biopsies and enables real-time monitoring of the tumor's genetic profile, facilitating personalized treatment decisions. Furthermore, Liquid Trace liquid biopsy can be used as a tissue-informed test for monitoring response and measurable residual disease with high sensitivity but at the same time detecting clonal evolution of new mutations.
Download Case Study Brochure (PDF)
References
- Albitar M., Zhang H., Charifa A., Ip A., De Dios I, Ma W., McCloskey J.K., Donato M, DiCapua Siegel D.S., Waintraub S.E,, Gutierrez M., Pecora A.L., Goy A. Combining cell-free RNA (cfRNA) with cell-free total nucleic acid (cfTNA) as a new paradigm for liquid biopsy. DOI: 10.1200/JCO.2022.40.16_suppl.3048 Journal of Clinical Oncology 40, no. 16_suppl (June 01, 2022) 3048-3048.
- Liao H., Huang W., Pei W., Li H. Detection of ESR1 mutations based on liquid biopsy in estrogen receptor-positive metastatic breast cancer: clinical impacts and prospects. Front Oncol. 2020 Dec 15;10:587671. doi: 10.3389/fonc.2020.587671.
Case Study: CSF - GBM
Background
A 66-year-old patient presented with progressive neurologic decline, imaging revealing a diffusely infiltrative lesion in the right temporal lobe with radiographic features suspicious for high-grade glioma. Cerebrospinal fluid (CSF) liquid biopsy was performed for genomic characterization. Both cfDNA and RNA NGS were applied to detect somatic mutations, structural alterations, and viral RNA.
Discussion
The Role of CSF cfDNA and RNA NGS
Traditional classification of CNS (Central Nervous System) tumors relies on tissue biopsy, which may be limited by anatomical inaccessibility of certain lesions and risk of complications from repeat biopsies.
1. Non-invasive “liquid biopsy” of the CNS – captures tumor DNA/RNA shed into the cerebrospinal fluid.
2. Comprehensive profiling – allows simultaneous detection of:
• Somatic mutations (TP53, PIK3CA, ARID2, etc.)
• Copy number alterations (EGFR amplification, +7/−10)
3. RNA expression patterns and fusions (if present).
4. Improved sensitivity in heterogeneous tumors – avoids sampling bias from a single tissue block.
5. Diagnostic confirmation – in this case, the hallmark +7/−10 signature and EGFR amplification firmly established glioblastoma, IDH-wildtype, without requiring invasive re-biopsy.
6. Therapeutic stratification – cfDNA and RNA profiling identified PIK3CA and ARID2 mutations, opening potential targeted therapy avenues.
7. Dynamic monitoring – serial CSF sampling can track tumor evolution, therapy resistance, and recurrence in real time.
Conclusion
This case illustrates the diagnostic and therapeutic value of CSF cfDNA and cfRNA NGS in CNS tumors. Beyond confirming glioblastoma, IDH-wildtype, the analysis uncovered actionable alterations and highlighted the possibility of germline predisposition. As molecular diagnostics evolve, CSF NGS will increasingly complement or even substitute tissue biopsy, offering a safer, more comprehensive, and dynamic method for CNS tumor classification and management.
Download Case Study Brochure (PDF)
References
- Miller et al., Nat Med 2019 – Demonstrated that CSF-derived cfDNA sequencing provides a more accurate representation of the glioma genome compared to plasma cfDNA, and captures mutations missed by tissue biopsy due to spatial heterogeneity.
- Pan et al., Clin Cancer Res 2019 – Showed that CSF cfDNA profiling detects EGFR, TP53, and other hallmark mutations in glioblastoma, highlighting its role in diagnosis and therapy guidance.
- Pentsova et al., Nat Commun 2016 – Pioneering study proving that CSF liquid biopsy can identify clinically relevant mutations in brain tumors, laying the foundation for its clinical utility.
- Wang et al., JCO Precis Oncol 2021 – Highlighted the utility of cfRNA in CSF for detecting gene fusions and expression profiles, further enhancing CNS tumor classification.
- Peripheral blood: 10 mL in an EDTA tube is required.
Important: RNA stability is 48-72 hours from blood draw. DNA stability is 7 days from blood draw. Samples received beyond 72 hours may include only DNA results.
- CSF: 7-10 mL is optimal (5 mL minimum).
Important: Ship as soon as possible (overnight). Do not use collection devices with anti-coagulants. Clear tubes.
Specimen Preparation and Shipping Guidelines
Use the Liquid Trace Transport Kit
- Complete the requisition, making sure that all sections are completed in their entirety, including client information, patient information, specimen information, and test selection. Missing information may delay reporting of test results.
- Diagnosis/patient history is extremely important in rendering the correct interpretation of results and should also be filled out as completely as possible. A copy of a pathology report should be included.
- Ensure the specimen is labeled with patient name and number. A minimum of two patient identifiers is required for each specimen.
For blood samples:
- Ship using a cold pack. The cold pack should not directly contact the blood tube. Ship as soon as the sample is collected, with overnight delivery.
Important: RNA stability is 48-72 hours from blood draw. DNA stability is 7 days from blood draw. Samples received beyond 72 hours may include only DNA results.
Request Kits
Fill out the form below to request kits. Please refer to the Specimen Requirements page for more details.
*GTC will need to set you up in our system if this is your first order.

How to complete the Genomic Testing Cooperative requisition form
Download our
Test Requisition
Keep in mind that we do not accept blood samples directly from individuals. Talk with your M.D. to fill out the form for you.
1. Rogers A, Joe Y, Manshouri T, Dey A, Jilani I, Giles F, Estey E, Freireich E, Keating M, Kantarjian H, Albitar M. Relative increase in leukemia-specific DNA in peripheral blood plasma from patients with acute myeloid leukemia and myelodysplasia. Blood. 2004;103:2799–801. 2. Nakamura S, Yokoyama K, Shimizu E, Yusa N, Kondoh K, Ogawa M, Takei T, Kobayashi A, Ito M, Isobe M, Konuma T, Kato S, Kasajima R, Wada Y, Nagamura-Inoue T, Yamaguchi R, Takahashi S, Imoto S, Miyano S, Tojo A. Prognostic impact of circulating tumor DNA status post-allogeneic hematopoietic stem cell transplantation in AML and MDS. Blood. 2019 Jun 20;133(25):2682-2695. doi: 10.1182/blood-2018-10-880690. 3. Paul Yeh, Michael Dickinson, Sarah Ftouni, Tane Hunter, Devbarna Sinha, Stephen Q. Wong, Rishu Agarwal, Ravikiran Vedururu, Kenneth Doig, Chun Yew Fong, Piers Blombery, David Westerman, Mark A. Dawson and Sarah-Jane Dawson. Molecular disease monitoring using circulating tumor DNA in myelodysplastic syndromes. Blood. 2017;129(12):1685-1690. 4. Albitar A, Ma W, DeDios I, Estella J, Ahn I, Farooqui M, Wiestner A, Albitar M. Using high-sensitivity sequencing for the detection of mutations in BTK and PLCγ2 genes in cellular and cell-free DNA and correlation with progression in patients treated with BTK inhibitors. Oncotarget. 2017 Mar 14;8(11):17936-17944. doi: 10.18632/oncotarget.15316. PMID: 28212557 5. Albitar F, Ma W, Diep K, De Dios I, Agersborg S, Thangavelu M, Brodie S, Albitar M. Deep Sequencing of Cell-Free Peripheral Blood DNA as a Reliable Method for Confirming the Diagnosis of Myelodysplastic Syndrome. Genet Test Mol Biomarkers. 2016 Jul;20(7):341-5. doi: 10.1089/gtmb.2015.0278. PMID: 27248906 6. Aljurf M, Abalkhail H, Alseraihy A, Mohamed SY, Ayas M, Alsharif F, Alzahrani H, Al-Jefri A, Aldawsari G, Al-Ahmari A, Belgaumi AF, Walter CU, El-Solh H, Rasheed W, Albitar M. Chimerism Analysis of Cell-Free DNA in Patients Treated with Hematopoietic Stem Cell Transplantation May Predict Early Relapse in Patients with Hematologic Malignancies. Biotechnol Res Int. 2016;2016:8589270. doi: 10.1155/2016/8589270. 7. Ma W, Kantarjian H, Zhang X, Jilani I, Sheikholeslami MR, Donahue AC, Ravandi F, Estey E, O’Brien S, Keating M, Giles FJ, Albitar M. Detection of nucleophosmin gene mutations in plasma from patients with acute myeloid leukemia: clinical significance and implications. Cancer Biomark. 2009;5(1):51-8. doi: 10.3233/CBM-2009-0583. PMID: 19242062 8. Yeh CH, Tseng R, Albitar M. Plasma-based detection of clonality in lymphoid malignancies. Eur J Haematol. 2009 Jun;82(6):450-3. doi: 10.1111/j.1600-0609.2009.01231.x. PMID: 19187275. 9. Ma W, Kantarjian H, Zhang X, Sun W, Buller AM, Jilani I, Schwartz JG, Giles F, Albitar M. Higher detection rate of JAK2 mutation using plasma. Blood. 2008 Apr 1;111(7):3906-7. doi: 10.1182/blood-2008-02-139188. PMID: 18362222. 10. Giles FJ, Albitar M. Plasma-based testing as a new paradigm for clinical testing in hematologic diseases. Expert Rev Mol Diagn. 2007 Sep;7(5):615-23. Review. PMID: 17892367. 11. Ma W, Tseng R, Gorre M, Jilani I, Keating M, Kantarjian H, Cortes J, O’Brien S, Giles F, Albitar M. Plasma RNA as an alternative to cells for monitoring molecular response in patients with chronic myeloid leukemia. Haematologica. 2007 Feb;92(2):170-5. PMID: 17296565. 12. Ma W, Kantarjian H, Jilani I, Gorre M, Bhalla K, Ottmann O, Giles F, Albitar M. Heterogeneity in detecting Abl kinase mutations and better sensitivity using circulating plasma RNA. Leukemia. 2006 Nov;20(11):1989-91. Epub 2006 Aug 24. PMID: 16932346. 13. Ma W, Jilani I, Gorre M, Keating M, Chan H, Tseng R, Kantarjian H, O’Brien S, Giles FJ, Albitar M. Plasma as a source of mRNA for determining IgV(H) mutation status in patients with chronic lymphocytic leukaemia. Br J Haematol. 2006 Jun;133(6):690-2. 14. Jilani I, Estey E, Manshuri T, Caligiuri M, Keating M, Giles F, Thomas D, Kantarjian H, Albitar M. Better detection of FLT3 internal tandem duplication using peripheral blood plasma DNA. Leukemia. 2003 Jan;17(1):114-9. 15. Kurtz DM, Green MR, Bratman SV, Scherer F, Liu CL, Kunder CA, Takahashi K, Glover C, Keane C, Kihira S, Visser B, Callahan J, Kong KA, Faham M, Corbelli KS, Miklos D, Advani RH, Levy R, Hicks RJ, Hertzberg M, Ohgami RS, Gandhi MK, Diehn M, Alizadeh AA.Noninvasive monitoring of diffuse large B-cell lymphoma by immunoglobulin high-throughput sequencing. Blood. 2015 Jun 11;125(24):3679-87.