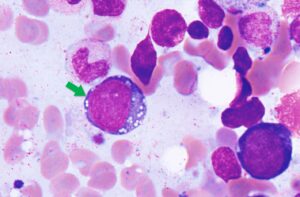
This Hematology Profile™ is designed to profile the molecular abnormalities in various hematologic neoplasms. The test analyzes the DNA of 302 genes that are associated with hematologic cancers including:
-Myelodysplastic syndrome (MDS)/Chronic myelomonocytic leukemia (CMML): The Hematology Profile is used for stratifying patients, determining prognosis and selecting therapy. This assay will not only determine the aggressiveness and prognosis of the MDS but will also determine if the patient has reactive cytopenia and will distinguish between CHIP (Clonal Hematopoiesis of Indeterminate Potential) or CCUS (Clonal Cytopenia of Unknown Significance) and MDS.
-VEXAS Syndrome: Recently described VEXAS (vacuoles, E1 enzyme, X-linked, autoinflammatory, somatic) is caused by mutations in the UBA1 gene. This is an adult-onset fatal disease that may present as myelodysplastic syndrome, aplastic anemia or multiple myeloma, but characterized by fevers, low white cell count, vacuoles in bone marrow cells, dysplastic bone marrow, pulmonary inflammation, chondritis, and vasculitis. Detecting the presence of mutations in the UBA1 gene is the only way for confirming the diagnosis of VEXAS syndrome.
-Acute Myeloid Leukemia (AML): The Hematology Profile helps in the diagnosis of AML and distinguish between De Novo AML vs secondary AML. It also helps in determining eligibility for treatment with FLT3 and IDH1/2 inhibitors. For a complete evaluation of translocations including all types of APL, Inv16, and t(8,21) and NUP98 translocations, we recommend ordering the GTC-Hematology PLUS panel.
-Myeloproliferative Neoplasms (MPN): This includes quantitative analysis of all exons of JAK2, CALR, and MPL.
-Lymphoma: Analysis of mutations reported in various types of lymphoma, including follicular, DLBCL, CLL, and T-cell lymphoma. However, for a complete evaluation of lymphoma including determining double hit lymphoma and diagnosis of ABC vs GBC, RNA adding the ordering GTC-Hematology PLUS panel is recommended.
-Multiple Myeloma: This Hematology Profile detects mutations in the coding sequence of genes frequently mutated in MM, but for a thorough evaluation, we recommend ordering GTC-Hematology PLUS to cover expression abnormalities and chromosomal translocations.
Clonal Hematopoiesis of Indeterminate Potential (CHIP): Distinguish CHIP from clinically active and relevant hematologic neoplasm based on an internally developed algorithm using variant allele frequency, chromosomal structural abnormalities, clinical and laboratory data and longitudinal data. This distinction is particularly important when evaluating minimal residual disease and in the presence of other neoplastic process.
Turn Around Time: 5-7 days
GTC uses AI in every step of our analysis and it makes a difference in helping make a new discovery daily that improves patient care.
Once the data is offloaded from the sequencer, our AI:
- Assists with mutation analysis, identifying non-mutations and artifacts
- Compares various data sets to explore disease biology
- Provides support for clinical decision-making and classification of the disease
- It helps with matching patients to therapeutics and presents clinical trial options
- Aggregates data for report generation and simplifies the results so they are easily understood
Case Study: VEXAS Syndrome
Background
VEXAS syndrome is a disease with inflammatory and hematologic (blood) manifestations. The syndrome is caused by mutations in the UBA1 gene affecting the Met41 residue of the protein and resulting in decreased cellular ubiquitylation activity and hyperinflammation. This is an adults-onset fatal disease that may present as myelodysplastic syndrome, aplastic anemia or multiple myeloma, but characterized by fevers, low white cell count, vacuoles in bone marrow cells, dysplastic bone marrow, pulmonary inflammation, chondritis, and vasculitis. Detecting UAB1 gene mutations in the is the only way for confirming the diagnosis of this disease.
Clinical History
68-year-old male
With low-grade myelodysplastic syndrome and inflammatory manifestations including arthritis, chondritis or other autoimmune disorders
Molecular Profiling Findings
Mutations in UBA1, DNMT3A, and TET2 genes
Discussion
The presence of DNMT3A, and TET2 suggest early low-grade myelodysplastic syndrome (MDS). And the detection of UBA1 mutation along with the patient’s symptoms confirmed the diagnosis of VEXAS. This patient was undiagnosed for long period of time and became transfusion dependent. By using any of our hematology DNA, DNA+RNA or cfDNA NGS panels; we are able to accurately diagnose VEXAS syndrome. This patient and most of the patients who are currently diagnosed with VEXAS have had numerous tests and tried multiple treatments. VEXAS should be considered in patients with systemic autoinflammatory disorders as well as patients with clinical presentation of myelodysplastic syndrome.
Download PDF (VEXAS case study brochure)
References
1. David B. Beck, M.D., et al. Somatic Mutations in UBA1 and Severe Adult-Onset Autoinflammatory Disease N Engl J Med 2020; 383:2628-2638, DOI: 10.1056/NEJMoa2026834
2. James A. Poulter, et al. Novel somatic mutations in UBA1 as a cause of VEXAS syndrome. Blood (2021) 137 (26): 3676–3681.
3. Marcela A. Ferrada, et al. Somatic Mutations in UBA1 Define a Distinct Subset of Relapsing Polychondritis Patients With VEXAS, Arithritis & Rheumatology, doi.org/10.1002/art.41743.
- Bone marrow: 2 mL. EDTA tube is preferred.
- Peripheral blood: 5 mL. EDTA tube is preferred.
- Fresh Tissue
- FFPE: 1 H&E slide and 6-10 unstained slides, 5-7 microns of BM clot or tissue fixed with 10% NBF fixative. Alternatively, the FFPE block of the BM clot can be sent for sectioning in our Lab.
Specimen Preparation and Shipping Guidelines
Use the Hematology Transport Kit or Solid Tumor Transport Kit (depending on the sample type)
- Complete Requisition, making sure all sections are completed in their entirety including client information, patient Information, specimen Information and test Selection. Missing information may delay reporting of test results.
- Diagnosis/patient history is extremely important in rendering the correct interpretation of results and should also be filled out as completely as possible. A copy of a Path report should be included.
- Ensure the specimen is labeled with patient name and number. A minimum of two patient identifiers is required for each specimen
For FFPE samples:
- If sending FFPE Block-Insert up to 6 blocks into plastic block tray provided. Insert block tray into foam insert in the transport box.
- If sending slides-Insert slides into plastic slide holders provided. Insert the slide holders into the foam insert in the transport box. The container will hold up to 6 maximum slide holders.
- Place folded test requisition(s) and/or manifest(s) if ordered electronically into the transport kit.
- Close box and tuck tabs into place. No tape necessary.
For blood samples:
- Ship using a cold pack. The cold pack should not directly contact the blood tube. Ship as soon as sample collected with overnight delivery.
Request Kits
Fill out the form below to request kits. Please refer to the Specimen Requirements page for more details.
*GTC will need to set you up in our system if this is your first order.
How to complete the Genomic Testing Cooperative requisition form.
Download our
Test Requisition
Keep in mind that we do not accept blood samples directly from individuals. Talk with your M.D. to fill out the form for you.
Do You Want to Download the Sample Report?